Building a RAG App
Building a RAG Application so I Know What a RAG Application Is
I like staying on top of current tech trends, and one thing that is very popular right now is retrieval-augmented generation (RAG) applications. That sounds intimidating, but in trying to build one I found, like most tech hype, that they are simultaneously very cool and very boring. Here I’ll describe my motivation, implementation, and future plans for a small RAG application I call LabLedger.
Endless Experiments with Poor Commits
I work mostly solo, meaning I’m my own manager for many of my daily coding tasks. In an effort to be a more competent computational biologist, I try and lean on the phrase “commit early, commit often”. This is also useful because I don’t really have a lab notebook. So if I go away for a long time, I like to be able to return and know what in the world I was working on.
A RAG application is a great opportunity to address this, and even comes with some unexpected uses. The main goal, however, will be to automate an “expert” code commit review and summary for each week. This can remind me of what I’ve done, where I should go next, and how I can make better commits.
Here’s a basic overview of the workflow for LabLedger from ingestion to LLM review:
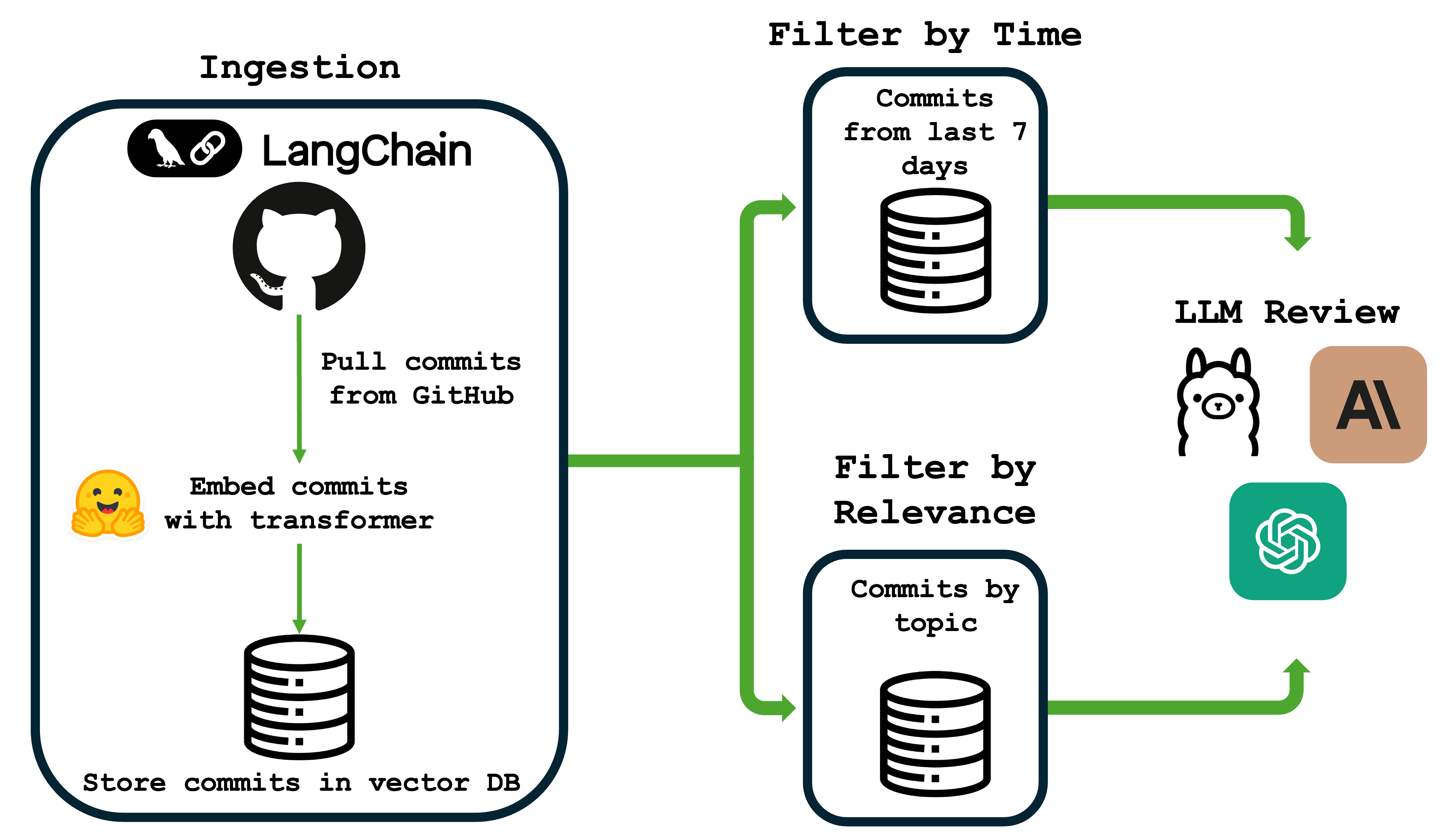
What is a RAG Application?
The primary goal of our RAG application will be to provide an LLM with focused information, so that it can cite things (such as our commits) without hallucination. It will:
- Retrieve our commits from a vector database.
- Augment an LLM prompt by injecting our commits into a custom “commit review” prompt.
- Generate a summary and review of our commits.
It’s actually that simple, and most of the work is in orchestrating the systems, not writing anything clever.
Setting Up a Vector Database With ChromaDB
A vector database will be a way to take embeddings of our commits, which are essentially numerical representations of each message meant to capture the meaning within context of the text. This has several advantages. Most importantly, this will allow us to perform semantic search. Semantic search means that we can feed in a prompt, such as “cell type prediction logic”. We embed this new message, then search across the vector database for n similar commits. For example, if I search the commits for “RNA” in our analysis repo for a submitted paper (https://github.com/brocklab/clusterCleaver-analysis) focusing on biomarker separation, we find these are the top 5 commits:
Commit: Output optimal genes
Commit: Processed data, initial surface gene search
Commit: Applying new separation across all genes
Commit: Removing outliers with DE genes
Commit: sc2markerComparison, added 436 umap
First, please note how bad these commits are. But secondly, notice how none of them have “RNA” in the commit but our outputs still have to do with genes. This is a powerful way for us to now review specific parts of our commit history based on the project.
Putting it Together With an LLM
While I’m probably way more interested in the actual embeddings and text similarity search, this is a good opportunity to get some focused feedback. Let’s ask Claude to rate these commits, provide a summary, and give some advice.
### Standup Summary
The team has made significant progress on RNA sequencing analysis, focusing on three main areas:
1. **10x Genomics Data Processing Pipeline**: Established initial processing workflows for 10x scRNA-seq data with both Python and R implementations, including data ingestion, quality control, and preliminary analysis.
2. **Optimal Gene Separation & Surface Marker Discovery**: Developed and optimized algorithms for identifying optimal gene separations across cell populations, with major performance improvements through vectorized Wasserstein distance calculations. Applied this methodology to identify surface markers and differentially expressed genes across multiple cell lines (HCC1806, HCC383, BT474, MDA-MB-231).
3. **Quality Control & Outlier Detection**: Implemented outlier removal strategies using differentially expressed (DE) genes, tested imputation effects on data quality, and performed cell cycle adjustments to reduce technical confounders in the analysis.
Additional work includes benchmarking against sc2marker tool, UMAP visualizations for cluster identification, and validation across multiple datasets including spleen tissue data.
### Commit Quality Audit
- **"Applying new separation across all genes"**: 6/10 - Vague commit message doesn't specify what "new separation" means or what the expected outcome is.
- **"Output optimal genes"**: 5/10 - Too generic; should specify which dataset, what optimization criteria, and how many genes were identified.
- **"Removing outliers with DE genes"**: 7/10 - Clear intent but lacks specifics on the DE threshold used or how many outliers were removed.
- **"Processed data, initial surface gene search"**: 6/10 - Combines two distinct activities (data processing and gene search) that should be separate commits.
- **"sc2markerComparison, added 436 umap"**: 4/10 - Cryptic message; "436" is unclear, and combining benchmarking with visualization is poor commit hygiene.
- **"Sorting out by DE genes"**: 3/10 - "Sorting out" is ambiguous; unclear what action was taken or why.
...
### Senior Advice
**Implement a structured commit message convention immediately.** Your commit history shows a pattern of vague, action-oriented messages ("Adjusted," "Made," "Testing") that provide minimal context for future debugging or onboarding. Adopt a format like: `[Component] Action: Specific change (Impact/Reason)`. For example, instead of "Adjusted UMAP," write: `[Visualization] Refactor UMAP: Updated n_neighbors=15 to reduce overclustering in HCC1806 dataset`. This is critical for RNA-seq work where parameter choices significantly impact biological interpretation. Additionally, your commits frequently bundle multiple concerns (data processing + analysis, benchmarking + visualization) - enforce atomic commits that change one logical thing at a time. This will dramatically improve your ability to bisect bugs and understand the evolution of your analysis pipeline.
I know it seems like a lot, but this has the power to completely alter my workflow by providing automated, focused feedback. We can also tamp down the number of commits (this cost me a whole 2 cents!), or restrict them to the past week, and so on. We could also use a locally-hosted LLM in case you’re using some more sensitive data.
Conclusion
While it seemed intimidating at first, this was hugely informative that RAG applications are actually pretty basic. I know we’re living in the world of LLM-wrappers, but this enables us to better understand our commits in a targeted way while minimizing hallucination. It also helps me better understand how and why my commits are so terrible.
One other thing I’d like to note that I learned is that AI coding tools are becoming increasingly important for my understanding of new tools. The way I like to learn things is typically by coming up with an idea, finding an example, and hacking at it until I understand the code and it starts to do what I want it to do. This usually involved a lot of googling, time on forums and Stack Overflow, and some documentation. Now it’s a lot different. I think good developers and coders will need to increasingly read more documentation in combination with mandatory AI coding tools. Unfortunately, code snippets are far easier to come by now from AI than anywhere else. However, I once again found these lacking and had to go do a lot of documentation digging. Clearly I have plenty of things to work on, but this was an amazing project for its practicality, novelty, and overall learning experience.
If you’re interested in digging more into the code, check out my repo on github here.